2025-04-03 10:54:17
HLA/MHC (Human) —— The Human MHC is Called HLA
HLA (human leucocyte antigen, human white blood cell antigen) is a group of highly polymorphic genes. The products encoded by the human HLA genes are called MHC (major histocompatibility complex, major histocompatibility complex), which are distributed on the surface of different types of cells. They not only determine the host's tissue compatibility but also determine the individual's ability to produce immune responses to different antigens. HLA is located on a polymorphic region of the short arm of human chromosome 6, which is about 3.6M in length and is involved in immune responses. The functions of HLA are twofold: one is tissue-specific antigens, and the other is antigen presentation.
The HLA system has high genetic polymorphism and is the main component of specific immune recognition and immune response within the body. Polymorphism (polymorphism) refers to the existence of multiple alleles (allele) at a gene locus. It is a concept of a population, indicating that there are differences in the allelic possession status among different individuals in a population. (Polygeny focuses on the changes at the HLA gene locus in the same individual, while polymorphism refers to the changes in the number of alleles at each locus in the population.)
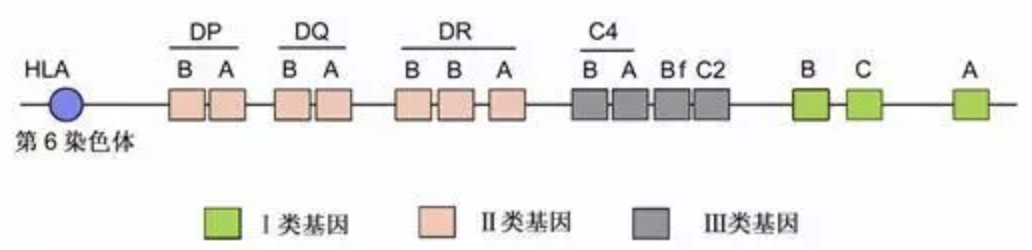
Figure: Structure of
The HLA complex is composed of many genes and can be roughly divided into three categories: Class I, Class II, and Class III. The two main types of HLA molecules involved in antigen presentation and related immune responses are Class I and Class II molecules. HLA-Class I includes HLA-A, -B, -C; HLA-Class II includes HLA-DR, -DQ, -DP. For a gene locus, an individual can have at most two alleles, one from each homologous chromosome inherited from the parents. Therefore, the surface of an individual's immune cells usually can detect 12 types of HLA Class I and Class II allelic molecules, i.e., HLA antigens, from both parents.
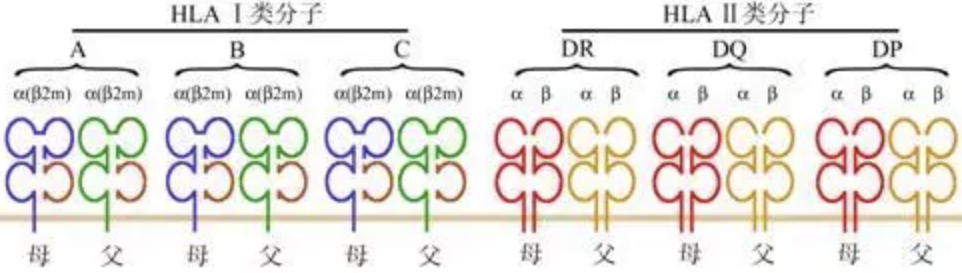
Figure: Co-dominant expression of 6 pairs of 12 classical Class I and Class II HLA molecules on the cell surface
HLA genes follow the laws of Mendelian genetics. The antigens encoded by them are co-dominantly expressed on the cell surface, meaning that both alleles at the corresponding loci on the homologous chromosomes can be expressed. Since the MHC genes are located on the same chromosome, the genotype combination at the polygenic loci is relatively stable, and recombination between homologous chromosomes rarely occurs. This constitutes the genetic feature of haplotype (HAPLOTYPE, i.e., a special combination of a series of alleles closely linked on the same chromosome).
Germ-line and Somatic HLA Testing
HLA testing is divided into two levels. One is the germline HLA genotyping of patients, which aims to examine the diversity of patients' HLA. The other level is to see whether the HLA genes in tumor cells have undergone loss of heterozygosity (LOH) due to immune escape. The former tests normal cells of patients, such as DNA extracted from peripheral blood leukocytes, while the latter tests DNA extracted from tumor tissues. HLA genotyping is the basis for the analysis of HLA heterozygosity.
|
HLA Testing Type |
Detection Content |
Sample Source |
Heritable |
|
Germline |
HLA genetic diversity (HLA typing) |
Normal cells (blood leukocytes) |
Yes |
|
Somatic |
HLA LOH (somatic LOH) |
Tumor tissue |
No |
Heritable HLA Allele Typing — HLA Diversity
With the exception of identical twins, no two individuals share identical HLA profiles, earning HLA the designation of "biological ID card" in human physiology
1)A complete HLA allele designation comprises 8 digits + 1 letter with the following structure
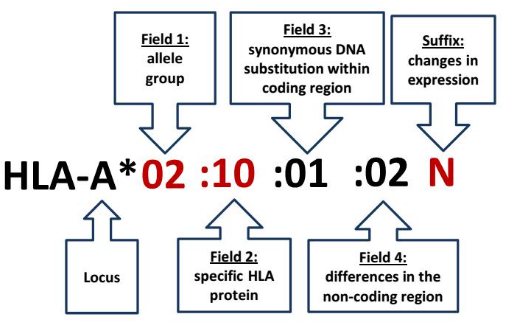
|
Component |
Explanation |
Example |
|
HLA |
Fixed prefix |
HLA |
|
A |
Gene locus (A/B/C/DR/DQ/DP) |
A |
|
* |
Separator |
* |
|
02 |
Allele group (serological specificity) |
02 |
|
10 |
Specific HLA protein variant |
10 |
|
01 |
Synonymous mutation in coding region |
01 |
|
02 |
Mutation in non-coding region |
02 |
|
N |
Expression status: |
N |
Typing Resolution Levels:
a) 2-digit: Allele group level
b) 4-digit: Specific protein level
c) 6-digit: Coding region polymorphism
d) 8-digit: Non-coding region variation
Classification of HLA Alleles
In 2007, the American Society for Histocompatibility and Immunogenetics (ASHI) established an ad hoc committee to propose a table of common and well-documented alleles (Common alleles and well documented alleles, CWD).
Strictly following the definition of CWD alleles in the latest international literature, HLA alleles are defined into three major categories: common alleles, well-documented alleles, and rare alleles.
Common alleles: Alleles with a frequency equal to or greater than 0.001 in the reference population.
Well-documented alleles: Alleles detected by PCR-SBT method in at least five independent unrelated individuals or in three independent unrelated individuals with specific haplotypes.
Rare alleles: All alleles other than common and well-documented alleles.
In CWD principle typing, common alleles and well-documented alleles are combined into CWD. When ambiguous allele combinations contain CWD alleles, further distinction is needed. However, when rare allele combinations appear, their clinical significance is limited and can be excluded.
The distribution of HLA genes has population characteristics, and the frequency of occurrence of HLA alleles varies among different populations and regions. The China Marrow Donor Program (CMDP) organized relevant experts in the domestic HLA field to formulate the CWD table for the Chinese population, which has important guiding significance for the routine gene typing work of HLA laboratories in China.
HLA Typing Methods
HLA typing is defined as the polymorphism of Class I and Class II loci. Serology is a traditional method for HLA typing. The rise of molecular detection methods has reduced the necessity of using sera containing all HLA antigen-specific antibodies for HLA typing. HLA gene typing technology is a molecular biology technique that has developed in recent years.
Sequence-based typing (SBT): This technology can be performed on multiple different platforms, one is based on Sanger DNA sequencing, and the other is based on NGS sequencing. The most common application of this method is to identify the HLA typing of stem cell transplant recipients and donors at the allelic level. An allele generally refers to a pair of genes located at the same position on a pair of homologous chromosomes that control relative traits and are unique nucleotide sequences on the gene. This method allows the confirmation of new allelic sequences.
The "gold standard" of HLA typing methods recommended by the World Health Organization is currently PCR-SBT (polymerase chain reaction-sequence specific primer), covering HLA-A, -B, -C, -DRB1, and -DQB1 loci, which are widely used for HLA gene typing of clinical transplant donors and recipients. However, due to the high polymorphism of HLA genes and the limitations of detection techniques, ambiguous allele typing results may occur in the typing of specific loci. Ambiguous genotype results refer to the existence of more than one possible combination of HLA alleles in the typing process of HLA genes.
Since NGS sequencing produces single-strand results, it helps to resolve the ambiguous assignment issues existing in classical double-strand sequencing. Although we have known about neoantigens for some time, it has only been in recent years, with the development of NGS technology and the improvement of accompanying bioinformatics analysis, that it has become possible to identify and characterize a large number of mutated neoantigens. HLA typing through next-generation sequencing data analysis can obtain polymorphisms ranging from single SNV genotypes to haplotype information. Through capture sequencing technology, the MHC region can be targeted for capture, resulting in higher sequencing depth and accuracy. The basic experimental workflow of NGS technology in HLA typing includes: template preparation, PCR system amplification, product normalization, sequencing reaction, and data analysis. NGS technology is highly throughput and accurate in HLA typing, and the cost of reagents is significantly reduced. However, it should be noted that there may be amplification imbalance and loss of HLA alleles during NGS detection.
Flowchart of HLA Typing by Next-Generation Sequencing:
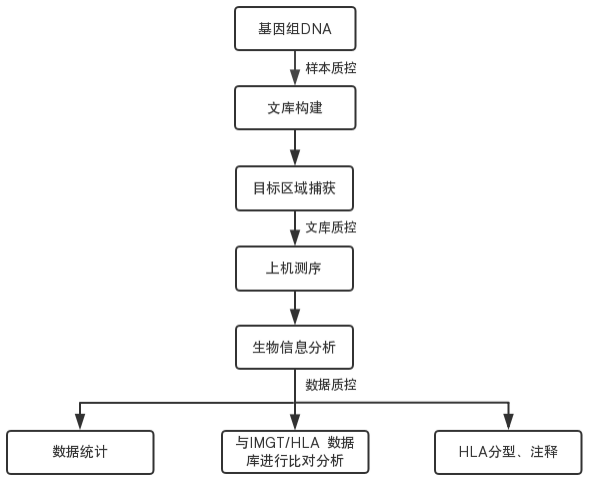
Loss of Heterozygosity (LOH) of HLA Genes —— Tumor Immune Escape
Loss of Heterozygosity (LOH): Under normal circumstances, a segment on an autosome will have one copy from the father and one from the mother. When LOH occurs, both chromosomes have the same segment from either the father or the mother, losing the corresponding segment from the other parent. This is called LOH.
Uniparental Disomy (UPD): A special form of LOH, where both chromosomes of a pair are from one parent (either father or mother), completely missing the corresponding chromosome from the other parent. The pathogenic potential of this variation is higher than that of LOH because it involves the loss of an entire chromosome.
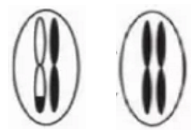
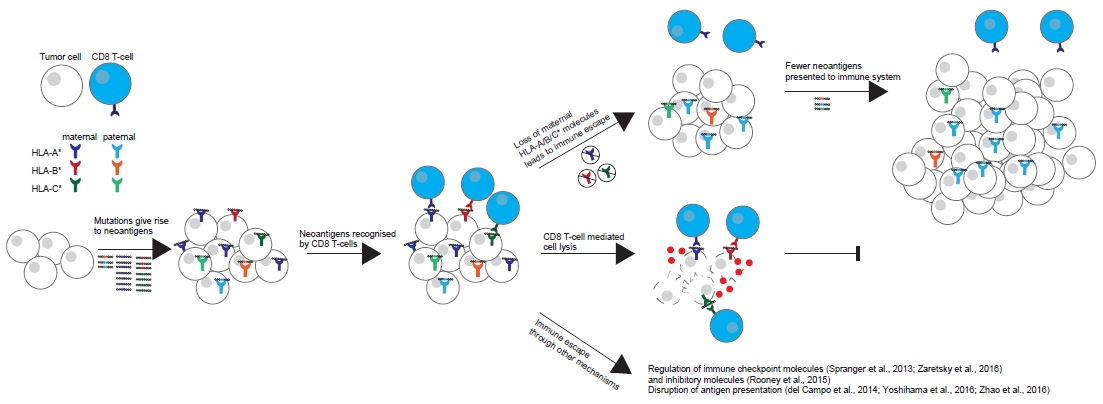
HLA gene LOH positivity indicates the presence of LOH in HLA genes, leading to impaired HLA function. Since the function of HLA is to present neoantigens to the cell surface for T cell recognition, partial or complete loss of function results in the inability to present antigens. The immune system cannot recognize the tumor, allowing the tumor to escape immune surveillance. This means that the patient may have a risk of resistance to immunotherapy drugs. HLA LOH is an immune escape mechanism adopted by tumor cells in response to immune system attacks, preventing recognition and killing by T cells, potentially leading to the accumulation of mutations in tumor cells. Immune escape due to HLA LOH is a significant barrier that immunotherapy needs to overcome.
Neoantigen (Neo-Antigen)
Neoantigens are abnormal proteins produced by cancer cell gene mutations that can activate the immune system. They carry specific amino acid sequence variations and can be presented by HLA molecules to induce T cell responses. Without amino acid sequence changes, these proteins would not be antigenic.。
Neoantigens are specific antigens located on the surface of tumor cells, characterized by tumor specificity. They are encoded by somatic mutations (somatic mutations) of oncogenes and contain antigenic peptides with mutated amino acids. These antigens do not exist in normal cells. Therefore, the immune system can recognize them as "non-self" antigens and trigger an immune response, unaffected by immune tolerance. This makes them ideal targets for immunotherapy. Neoantigens, derived from tumor-specific somatic mutations, exhibit individual differences and tumor specificity. They can be recognized by T cells and induce specific immune responses, holding great promise in the field of personalized immunotherapy.。
Tumor neoantigens, as targets for immunotherapy, have several major advantages: no central tolerance, strong affinity with major histocompatibility complex (MHC) molecules, reduced off-target side effects in normal tissues, and high safety.
When neoantigens are inside the cell, they are cleaved into peptides, which bind to Class I or Class II molecules. Then, HLA molecules present these peptides on the cell surface for recognition by T cell receptors. HLA Class I and Class II molecules present short peptides derived from pathogens (such as tumor neoantigens) to T cells to initiate adaptive immune responses. The fundamental reason why neoantigens are effective in immunotherapy is that they can activate immune cells to recognize and kill tumor cells.
Prediction of Tumor Neoantigens
Increasing evidence suggests that only a small fraction of non-synonymous mutations identified by tumor whole-exome sequencing (WES) can produce immunogenic neoantigens. The success of neoantigen identification largely depends on the identification of candidate neoantigens and the assessment of their immunogenicity.
Tumor neoantigen prediction involves two main key points: one is HLA typing of the patient's HLA molecules; the other is the identification and selection of suitable somatic mutations. Tumor neoantigen prediction generally requires precise HLA subtyping to the four-digit level.
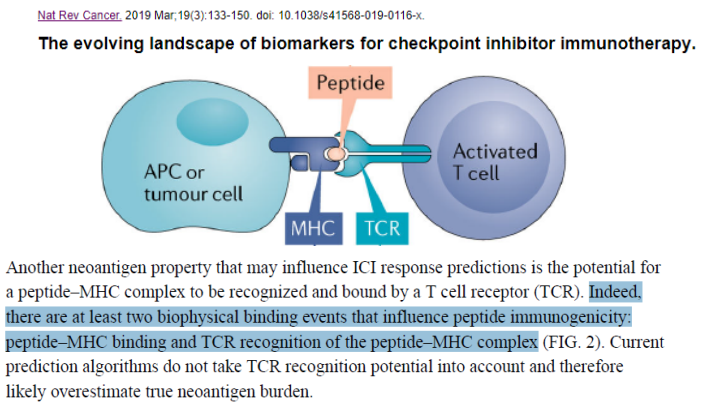
How to better identify immunogenic neoantigens from genomic data is one of the important issues to be solved at present. In fact, there are at least two bio-physical binding events that affect peptide immunogenicity: peptide-MHC binding and TCR recognition of the peptide-MHC complex. Current methods can be divided into two types: one is to predict the binding affinity of mutated peptides with HLA typing, and the other is to predict the potential of T cells' TCR to recognize neoantigens.
When selecting neoantigens, priority should be given to those with deep sequencing depth, high mutation frequency, strong mRNA expression signals, low affinity (half-maximal inhibitory concentration IC50) for HLA molecules (indicating stronger binding), and a small ratio of neoantigen affinity to that of the corresponding wild-type peptide (meaning that the neoantigen binds much stronger to HLA molecules than the wild-type peptide). In addition, the biological function of the mutated gene generating the neoantigen should also be considered, especially its association with the diseas under study.
The Three Pillars of Tumor Immunotherapy: Immune Checkpoint Inhibitors, Adoptive Cell Therapy, and Personalized Tumor Vaccines
Cancer immunotherapy is a treatment that uses immune cells to enhance their function to eliminate cancer. It can: help the immune system recognize and attack specific cancer cells; strengthen immune cells to destroy cancer; and provide additional substances to enhance the body's immune response.
The main immunotherapy technologies we usually refer to are as follows:
Immune checkpoint inhibitors: By inhibiting the immune escape of tumor cells, they mobilize the body's own immune system to eliminate tumors, represented by CTLA-4/PD-1/PD-L1 monoclonal antibodies.
Tumor ACT therapy (adoptive cell therapy) refers to the infusion of immune cells with antitumor activity that have been cultured and activated in vitro into patients. They directly kill tumor cells or activate the body's immune system to destroy tumor cells, ultimately achieving the goal of tumor treatment. Representative examples include CAR-T/TCR-T. Adoptive cells carrying tumor-specific antigens (tumor neoantigens/Neoantigen) can specifically activate T cells carrying Neoantigen to selectively kill tumor cells.
Personalized tumor vaccines refer to the injection of tumor neoantigen (neoantigen) peptides or RNA into patients' bodies to activate the immune system and achieve the effect of killing tumors. The breakthrough in personalized tumor vaccine technology lies in the development of neoantigen detection and prediction technologies to prepare neoantigen vaccines.
So far, clinical trial vaccines targeting neoantigens have been proven to be safe and well-tolerated, and personalized T cell therapies targeting neoantigens have shown antitumor responses.
Summary: Tumor immunotherapy based on neoantigens has the following advantages: (1) Neoantigens are only expressed in tumor cells and are non-heritable; (2) This therapy only activates the individual's own immune system and does not introduce viruses or other substances that did not previously exist in the individual, minimizing risks; (3) High targeting accuracy.
Note: TCR (T cell receptor) and CAR (chimeric antigen receptor) T cell therapies both involve modifying the patient's own T lymphocytes outside the body and then reinfusing them to kill tumors. However, their mechanisms of antigen recognition are completely different. TCRs use heterodimers composed of α and β peptide chains to recognize peptide fragments presented on the cell surface by MHC (major histocompatibility complex). In contrast, CARs use antibody fragments that can bind to specific antigens to recognize antigens on the surface of tumor cells, including tumor-specific antigens (TSA) and tumor-associated antigens (TAA). CAR-T cell therapy has achieved great development in the treatment of blood cancers, but the effectiveness of CAR-T therapy in solid tumors is not as optimistic. The main problem is that their target antigens are cell surface proteins, and the antigen expression of solid tumors is highly heterogeneous. Selecting suitable CAR-T cell targets in solid tumors is a challenge faced by researchers. In the treatment of solid tumors, TCR therapy may have advantages over CAR therapy.
Biomarkers for Immunotherapy Prognosis: Human Leukocyte Antigen (HLA) and Neoantigens Human HLA Gene Diversity
HLA Loss of Heterozygosity (LOH)
The higher the allelic heterozygosity of HLA genes, the better the prognosis for patients undergoing treatment. Patients with homozygous HLA alleles or those exhibiting LOH tend to respond poorly to immune checkpoint inhibitor therapy. LOH in HLA genes is an immune escape mechanism employed by cancers in response to strong selective pressures.
TMB in Combination with HLA LOH
Analyzing TMB together with HLA LOH can more accurately identify patients who are likely to benefit from treatment. Advanced tumors may opt for HLA LOH as a means of immune escape. Therefore, relying solely on TMB to guide treatment decisions could lead to ineffective immunotherapy in some patients.
Tumor Neoantigen Burden (TNB)
An elevated tumor mutation burden (TMB) increases the likelihood of generating neoantigens, thereby enhancing immunogenicity. The true tumor neoantigen burden (TNB) may have a stronger association with response to immune checkpoint inhibitors (ICI) compared to TMB.
Neo-Antigen (Neoantigen)
Generally speaking, the higher the total number of gene mutations in a patient's tumor tissue (i.e., the higher the TMB), the greater the number of neoantigens present. This increases the probability of specific recognition by T cell receptors (TCRs), thereby enhancing the immune-killing activity of neoantigen vaccines against tumors.
Note: TNB (Tumor Neoantigen Burden) is an indicator reflecting the total number of neoantigens in tumor cells, typically expressed as the number of tumor neoantigens per million bases (Mb) in the tumor genome region. TNB may emerge as a biomarker in cancer immunotherapy, driving the development of new treatment methods that selectively enhance T cell reactivity against these antigens.
References:
[1] 中华骨髓库CWD表2.2版本.
[2] 陈碧清,孔祥图,徐祖琼,代兴斌,于菊华,朱学军.原发性浆细胞白血病肿瘤新生抗原的筛查和预测[J].中国肿瘤生物治疗杂志,2019,26(07):730-735.
[3] Havel, J. J., D. Chowell and T. A. Chan (2019). "The evolving landscape of biomarkers for checkpoint inhibitor immunotherapy." 19(3): 133-150.
[4] McGranahan, N., et al. (2017)."Allele-Specific HLA Loss and Immune Escape in Lung Cancer Evolution." Cell 171(6):1259-1271 e1211.
[5] Chowell, D. and L. G. T. Morris (2018). "Patient HLA class I genotype influences cancer response to checkpoint blockade immunotherapy." 359(6375): 582-587.




